Conditional VAE#
Recall that our VAE setup:
The encoder \(f_{\phi}(x)\) takes a batch of images, (64, 28, 28), and returns \(\mu_z\) and \(\log \sigma^2_z\), each of (64, 2), s.t.
The decoder \(f_\theta(z)\) takes a batch of \(\hat{z} = \mu_z + \sigma_z \cdot \epsilon \sim q(z \mid x)\), and returns the mean of the reconstructed image \(\mu_x\) (64, 28, 28)
where we assumed \(\sigma^2_x=0.5\), s.t.
With cVAE, we simply make the class labels as additional inputs to the encoder and decoder networks.
s.t. we obtain posterior \(q_\phi(z\mid x, y)\) and \(p_\theta(x \mid z, y)\)
import os
os.environ['KERAS_BACKEND'] = 'torch'
import keras
import tensorflow as tf
import torch
import torch.nn.functional as F
from keras import ops
import numpy as np
import matplotlib.pyplot as plt
from keras.models import Sequential, Model
from keras.layers import Input, Dense, Flatten, Concatenate
Encoder#
The encoder \(f_{\phi}(x, y)\) takes a batch of images, (64, 28, 28), and one-hot labels (64, 10)
returns \(\mu_z\) and \(\log \sigma^2_z\), each of (64, 2), s.t.
image_input = Input(shape=(28, 28), name='image')
label_input = Input(shape=(10,), name='label')
latent_dim = 2
h = Flatten()(image_input)
h = Concatenate(axis=-1)([h, label_input])
h = Dense(512, activation='relu')(h)
h = Dense(32, activation='relu')(h)
h = Dense(2 * latent_dim)(h)
z_mean, z_log_var = ops.split(h, 2, axis=-1)
encoder = Model(inputs=[image_input, label_input], outputs=[z_mean, z_log_var], name='encoder')
encoder.summary()
Model: "encoder"
┏━━━━━━━━━━━━━━━━━━━━━┳━━━━━━━━━━━━━━━━━━━┳━━━━━━━━━━━━┳━━━━━━━━━━━━━━━━━━━┓ ┃ Layer (type) ┃ Output Shape ┃ Param # ┃ Connected to ┃ ┡━━━━━━━━━━━━━━━━━━━━━╇━━━━━━━━━━━━━━━━━━━╇━━━━━━━━━━━━╇━━━━━━━━━━━━━━━━━━━┩ │ image (InputLayer) │ (None, 28, 28) │ 0 │ - │ ├─────────────────────┼───────────────────┼────────────┼───────────────────┤ │ flatten (Flatten) │ (None, 784) │ 0 │ image[0][0] │ ├─────────────────────┼───────────────────┼────────────┼───────────────────┤ │ label (InputLayer) │ (None, 10) │ 0 │ - │ ├─────────────────────┼───────────────────┼────────────┼───────────────────┤ │ concatenate │ (None, 794) │ 0 │ flatten[0][0], │ │ (Concatenate) │ │ │ label[0][0] │ ├─────────────────────┼───────────────────┼────────────┼───────────────────┤ │ dense (Dense) │ (None, 512) │ 407,040 │ concatenate[0][0] │ ├─────────────────────┼───────────────────┼────────────┼───────────────────┤ │ dense_1 (Dense) │ (None, 32) │ 16,416 │ dense[0][0] │ ├─────────────────────┼───────────────────┼────────────┼───────────────────┤ │ dense_2 (Dense) │ (None, 4) │ 132 │ dense_1[0][0] │ ├─────────────────────┼───────────────────┼────────────┼───────────────────┤ │ split (Split) │ [(None, 2), │ 0 │ dense_2[0][0] │ │ │ (None, 2)] │ │ │ └─────────────────────┴───────────────────┴────────────┴───────────────────┘
Total params: 423,588 (1.62 MB)
Trainable params: 423,588 (1.62 MB)
Non-trainable params: 0 (0.00 B)
Decoder#
The decoder \(f_{\theta}(z, y)\) takes a batch of \(z\) (64, 2), and the corresponding one-hot labels \(y\) (64, 10), where \(y\) should correspond to the \(x\) that generates \(z\) in \(q(z \mid x)\)
returns the mean of the reconstructed image \(\mu_x\) (64, 28, 28)
where we assumed \(\sigma^2_x=0.5\), s.t.
from keras.layers import Reshape
z_input = Input(shape=(latent_dim,), name='z')
h = Concatenate(axis=-1)([z_input, label_input])
h = Dense(32, activation='relu')(h)
h = Dense(512, activation='relu')(h)
h = Dense(28 * 28)(h)
h = Reshape((28, 28))(h)
x_mean = ops.sigmoid(h)
decoder = Model(inputs=[z_input, label_input], outputs=x_mean, name='decoder')
decoder.summary()
Model: "decoder"
┏━━━━━━━━━━━━━━━━━━━━━┳━━━━━━━━━━━━━━━━━━━┳━━━━━━━━━━━━┳━━━━━━━━━━━━━━━━━━━┓ ┃ Layer (type) ┃ Output Shape ┃ Param # ┃ Connected to ┃ ┡━━━━━━━━━━━━━━━━━━━━━╇━━━━━━━━━━━━━━━━━━━╇━━━━━━━━━━━━╇━━━━━━━━━━━━━━━━━━━┩ │ z (InputLayer) │ (None, 2) │ 0 │ - │ ├─────────────────────┼───────────────────┼────────────┼───────────────────┤ │ label (InputLayer) │ (None, 10) │ 0 │ - │ ├─────────────────────┼───────────────────┼────────────┼───────────────────┤ │ concatenate_1 │ (None, 12) │ 0 │ z[0][0], │ │ (Concatenate) │ │ │ label[0][0] │ ├─────────────────────┼───────────────────┼────────────┼───────────────────┤ │ dense_3 (Dense) │ (None, 32) │ 416 │ concatenate_1[0]… │ ├─────────────────────┼───────────────────┼────────────┼───────────────────┤ │ dense_4 (Dense) │ (None, 512) │ 16,896 │ dense_3[0][0] │ ├─────────────────────┼───────────────────┼────────────┼───────────────────┤ │ dense_5 (Dense) │ (None, 784) │ 402,192 │ dense_4[0][0] │ ├─────────────────────┼───────────────────┼────────────┼───────────────────┤ │ reshape (Reshape) │ (None, 28, 28) │ 0 │ dense_5[0][0] │ ├─────────────────────┼───────────────────┼────────────┼───────────────────┤ │ sigmoid (Sigmoid) │ (None, 28, 28) │ 0 │ reshape[0][0] │ └─────────────────────┴───────────────────┴────────────┴───────────────────┘
Total params: 419,504 (1.60 MB)
Trainable params: 419,504 (1.60 MB)
Non-trainable params: 0 (0.00 B)
Full setup#
Preprocessing#
from keras.datasets import mnist
from torch.utils.data import TensorDataset, DataLoader
(x_train, y_train), (x_val, y_val) = mnist.load_data()
def preprocess(image, label):
return ops.cast(image, 'float32') / 255., ops.one_hot(label, 10)
x_train, y_train = preprocess(x_train, y_train)
x_val, y_val = preprocess(x_val, y_val)
train_ds = TensorDataset(x_train, y_train)
val_ds = TensorDataset(x_val, y_val)
train_loader = DataLoader(train_ds, batch_size=64, shuffle=True)
val_loader = DataLoader(val_ds, batch_size=64)
CVAE#
from keras.models import Model
from keras.metrics import Mean
import numpy as np
class CVAE(Model):
def __init__(self, encoder, decoder, **kwargs):
super().__init__(**kwargs)
self.encoder = encoder
self.decoder = decoder
self.loss_metric = Mean(name='loss')
self.nll_metric = Mean(name='nll')
self.kl_metric = Mean(name='kl')
self.pi = ops.array(np.pi)
def _get_losses(self, data):
image, label = data
z_mean, z_log_var = self.encoder([image, label])
batch_size, latent_dim = ops.shape(z_mean)[0], ops.shape(z_mean)[1]
epsilon = keras.random.normal((5, batch_size, latent_dim))
z_std = ops.exp(0.5 * z_log_var)
posterior_samples = z_mean + (z_std * epsilon) # (5, B, L)
log_Z = 0.5 * ops.log(2 * self.pi)
prior_log_prob = -0.5 * ops.square(posterior_samples) - log_Z
prior_log_prob = ops.mean(ops.sum(prior_log_prob, axis=-1))
posterior_log_prob = -0.5 * ops.square((posterior_samples - z_mean) / z_std) - ops.log(z_std) - log_Z
posterior_log_prob = ops.mean(ops.sum(posterior_log_prob, axis=-1))
kl_loss = posterior_log_prob - prior_log_prob
epsilon = keras.random.normal(ops.shape(z_mean))
z_sample = posterior_samples[0]
x_mean = self.decoder([z_sample, label])
nll_loss = 0.5 * ops.square((image - x_mean) / 0.5) + ops.log(0.5) + log_Z
nll_loss = ops.mean(ops.sum(nll_loss, axis=[-1, -2]))
loss = kl_loss + nll_loss
return loss, kl_loss, nll_loss
def call(self, inputs):
if isinstance(inputs, tuple):
image, label = inputs
else:
print(inputs.shape)
raise ValueError(f"Expected (image, label) tuple, got {type(inputs)}")
z_mean, z_log_var = self.encoder(inputs)
epsilon = keras.random.normal(ops.shape(z_mean))
z_std = ops.exp(0.5 * z_log_var)
z_sample = z_mean + (z_std * epsilon)
return self.decoder([z_sample, label])
def train_step(self, data):
backend = keras.config.backend()
if backend == 'tensorflow':
with tf.GradientTape() as tape:
loss, kl_loss, nll_loss = self._get_losses(data)
grads = tape.gradient(loss, self.trainable_weights)
self.optimizer.apply_gradients(zip(grads, self.trainable_weights))
elif backend == 'torch':
self.zero_grad()
loss, kl_loss, nll_loss = self._get_losses(data)
loss.backward()
gradients = [v.value.grad for v in self.trainable_weights]
with torch.no_grad():
self.optimizer.apply(gradients, self.trainable_weights)
else:
raise NotImplementedError(f"Unsupported backend: {backend}")
self.loss_metric.update_state(loss)
self.nll_metric.update_state(nll_loss)
self.kl_metric.update_state(kl_loss)
return {m.name: m.result() for m in self.metrics}
def test_step(self, data):
loss, kl_loss, nll_loss = self._get_losses(data)
self.loss_metric.update_state(loss)
self.nll_metric.update_state(nll_loss)
self.kl_metric.update_state(kl_loss)
return {m.name: m.result() for m in self.metrics}
@property
def metrics(self):
return [self.loss_metric, self.nll_metric, self.kl_metric]
Training#
cvae = CVAE(encoder, decoder, name='cvae')
cvae.compile(optimizer='adam')
# Manually call the model with a batch of data to build it
for batch in train_loader:
cvae((batch[0], batch[1])) # explicitly pass (image, label)
break
cvae.fit(train_loader, validation_data=val_loader, epochs=10)
Epoch 1/10
938/938 ━━━━━━━━━━━━━━━━━━━━ 13s 14ms/step - kl: 4.9593 - loss: 277.4842 - nll: 272.5248 - val_kl: 3.5252 - val_loss: 243.7113 - val_nll: 240.1861
Epoch 2/10
938/938 ━━━━━━━━━━━━━━━━━━━━ 13s 14ms/step - kl: 3.4908 - loss: 242.5442 - nll: 239.0534 - val_kl: 3.6831 - val_loss: 239.3537 - val_nll: 235.6707
Epoch 3/10
938/938 ━━━━━━━━━━━━━━━━━━━━ 12s 13ms/step - kl: 3.7619 - loss: 238.8095 - nll: 235.0477 - val_kl: 3.7457 - val_loss: 237.2413 - val_nll: 233.4957
Epoch 4/10
938/938 ━━━━━━━━━━━━━━━━━━━━ 13s 14ms/step - kl: 3.8970 - loss: 236.9694 - nll: 233.0724 - val_kl: 3.8853 - val_loss: 235.9740 - val_nll: 232.0887
Epoch 5/10
938/938 ━━━━━━━━━━━━━━━━━━━━ 13s 14ms/step - kl: 4.0104 - loss: 235.7918 - nll: 231.7813 - val_kl: 3.9562 - val_loss: 235.2438 - val_nll: 231.2875
Epoch 6/10
938/938 ━━━━━━━━━━━━━━━━━━━━ 13s 14ms/step - kl: 4.0717 - loss: 235.2486 - nll: 231.1769 - val_kl: 4.1178 - val_loss: 234.9267 - val_nll: 230.8090
Epoch 7/10
938/938 ━━━━━━━━━━━━━━━━━━━━ 13s 13ms/step - kl: 4.1322 - loss: 234.5730 - nll: 230.4409 - val_kl: 3.9452 - val_loss: 234.6644 - val_nll: 230.7191
Epoch 8/10
938/938 ━━━━━━━━━━━━━━━━━━━━ 13s 13ms/step - kl: 4.1654 - loss: 234.2233 - nll: 230.0579 - val_kl: 4.1858 - val_loss: 234.2850 - val_nll: 230.0992
Epoch 9/10
938/938 ━━━━━━━━━━━━━━━━━━━━ 13s 14ms/step - kl: 4.2182 - loss: 233.8293 - nll: 229.6111 - val_kl: 4.1584 - val_loss: 234.0365 - val_nll: 229.8781
Epoch 10/10
938/938 ━━━━━━━━━━━━━━━━━━━━ 13s 14ms/step - kl: 4.2465 - loss: 233.5610 - nll: 229.3145 - val_kl: 4.2682 - val_loss: 233.7383 - val_nll: 229.4701
<keras.src.callbacks.history.History at 0x7fb31bb63df0>
Visualization#
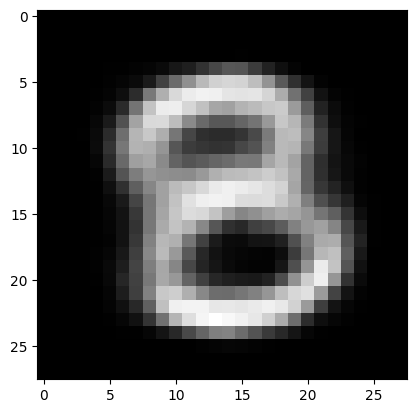
Generate \( \hat{z} \) from the prior \( \mathbb{N}(0, I) \), choose a digit \(c\) from 0-9, and simply input both to the decoder:
digit = 8
label = ops.one_hot([digit], 10)
sample = decoder([keras.random.normal((1, latent_dim)), label])[0]
sample = sample.detach().cpu().numpy()
plt.imshow(sample, cmap='gray')
<matplotlib.image.AxesImage at 0x7fb31bb4d4f0>
What if the z does not correspond to the y#
What if you give a random \( z \) and a label \( y \) to the decoder?
If \( z \sim p(z) = \mathcal{N}(0, I) \) (prior sampling):#
The decoder will try to produce an \( x \) that “looks like” class \( y \), as long as the decoder was trained well and the sampled \( z \) is within the region it saw during training.
If \( z \) is unrealistic, or corresponds to another class \( y' \):#
The decoder will still try to generate something that matches \( y \), but:
The output might look off or noisy.
You’re feeding it a latent that may not lie on the manifold of that class.
The decoder might “struggle” and output an ambiguous or nonsensical image.
The decoder is trained to map combinations of \( z \) and \( y \) to valid images.
But it only saw certain combinations during training: \( z \sim q(z|x, y) \), for real \( x \)‘s in class \( y \).
Say the encoder learns that \(z_1\) is the posterior mean to \(x_1, y_1\) in \(q(z | x, y)\)
If we feed \(z_1, y_2\) to the CVAE, or try to make an image of class \( y_2 \) using the latent from class \( y_1 \), it may or may not work well.
The model may be able to interpolate \( z \) between two real latent vectors of class \( y \), and still get reasonable outputs
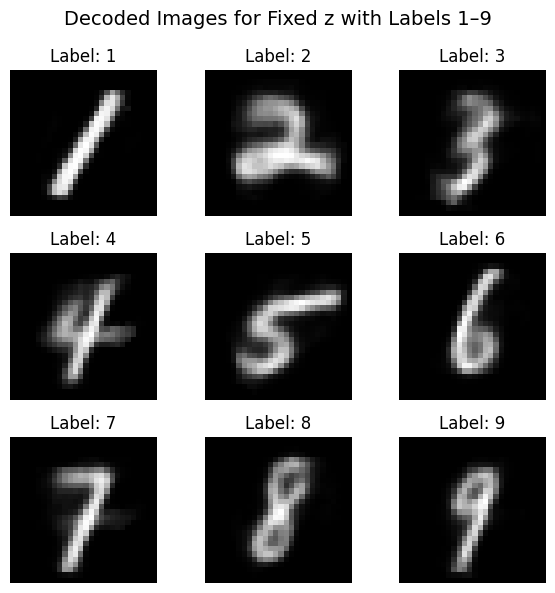
Here we try looping over a fixed \( z \) and change \( y \) across all classes:
“draw what this latent looks like as if it were each class”
z_rand = keras.random.normal((1, latent_dim))
fig, axs = plt.subplots(3, 3, figsize=(6, 6))
fig.suptitle("Decoded Images for Fixed z with Labels 1–9", fontsize=14)
for i, digit in enumerate(range(1, 10)):
label = ops.one_hot([digit], 10)
decoded = decoder([z_rand, label])[0]
image = decoded.detach().cpu().numpy()
ax = axs[i // 3, i % 3]
ax.imshow(image, cmap='gray')
ax.set_title(f"Label: {digit}")
ax.axis('off')
plt.tight_layout()
plt.subplots_adjust(top=0.88)
plt.show()

